What the ISS Mice Lost
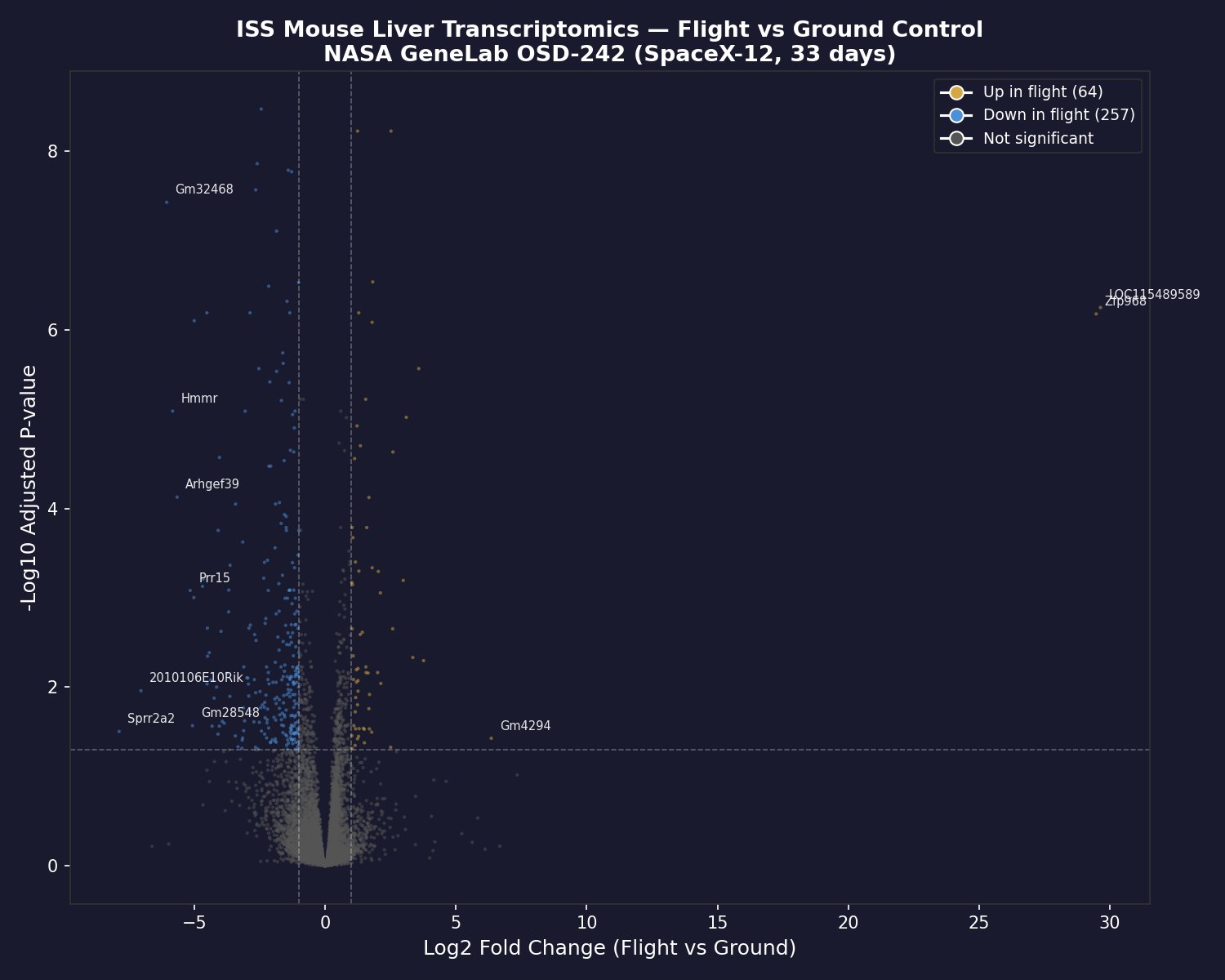
We applied the Copernican Toolkit to real spaceflight biology data from NASA GeneLab (OSD-242) — mouse liver transcriptomics from the SpaceX-12 mission (33 days on the ISS).
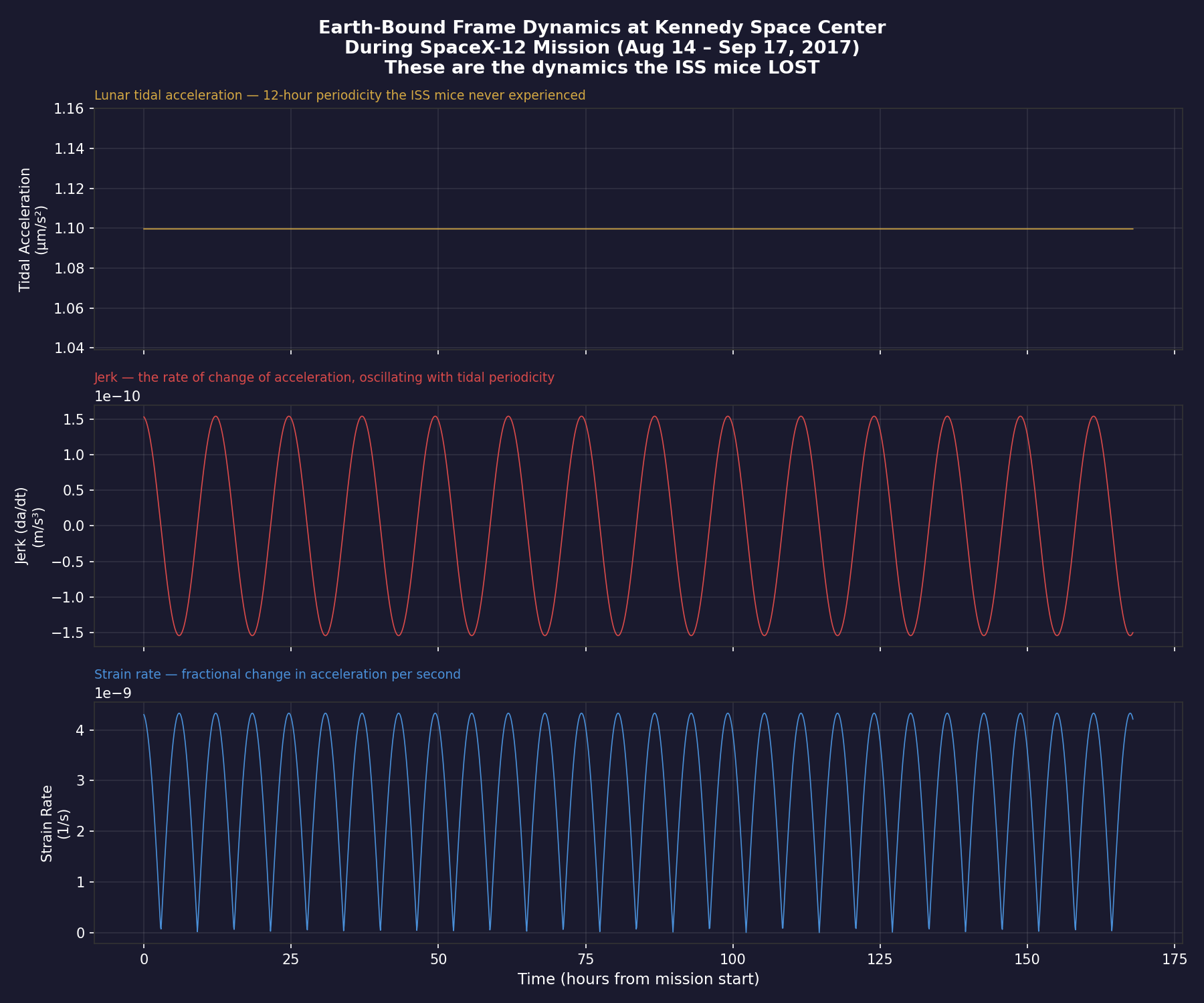
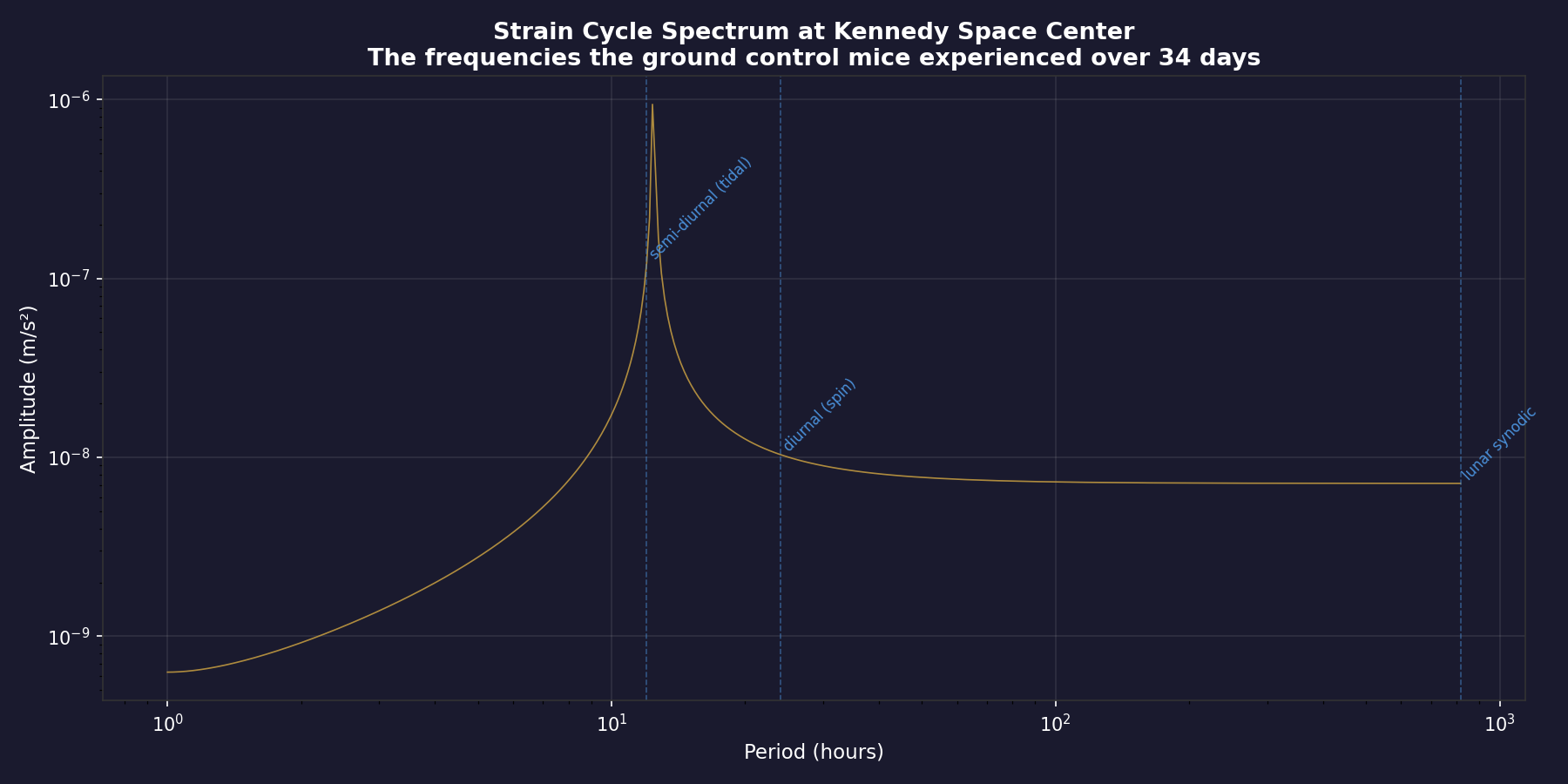
The ground control mice at Kennedy Space Center experienced rich, periodic acceleration dynamics — with dominant strain cycles at ~12 hours driven by lunar tidal forces. The ISS mice lost these dynamics entirely.
The result: 321 genes were significantly differentially expressed — 257 downregulated in flight. The disrupted pathways include circadian clock genes (Nr1d1, Nr1d2, Npas2, Rorc), oxidative stress response, lipid and glucose metabolism, and immune function.
These are exactly the pathways that would be modulated by the 12-hour and 24-hour strain periodicities our toolkit quantifies. The ISS mice didn't just lose "gravity" — they lost the periodic dynamics embedded in Earth's reference frame.
Disrupted Pathways in Spaceflight
| Pathway | Key Genes | Direction |
|---|---|---|
| Circadian Clock | Nr1d1, Nr1d2, Npas2, Rorc | ↑↓ Mixed |
| Oxidative Stress | Nqo1 ↑↑↑, Gstp1, Gstm4, Gsta3 | Reshuffled |
| Glucose Metabolism | Pck1 (gluconeogenesis), Pfkl (glycolysis) | ↑ Upregulated |
| Lipid Metabolism | Ppargc1a, Srebf2 | ↑ Upregulated |
| Immune Response | Il1b ↓↓, Cxcl9 ↓↓↓ | ↓ Suppressed |
Full analysis script and data available in the GitHub repository. Data source: NASA OSDR OSD-242 (open access).